MorphAEus
Unsupervised Outlier Detection
Common Assumption:
Auto-Encoders (AEs) trained on a data distribution (e.g., healthy chest X-rays) can only reconstruct in-distribution (ID) samples well, and yield high reconstruction error for out-of-distribution (OOD) samples:
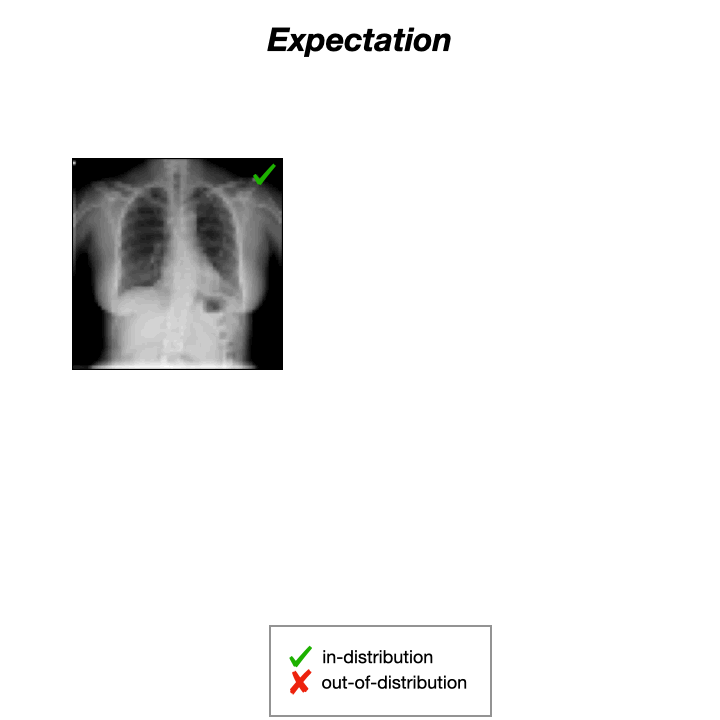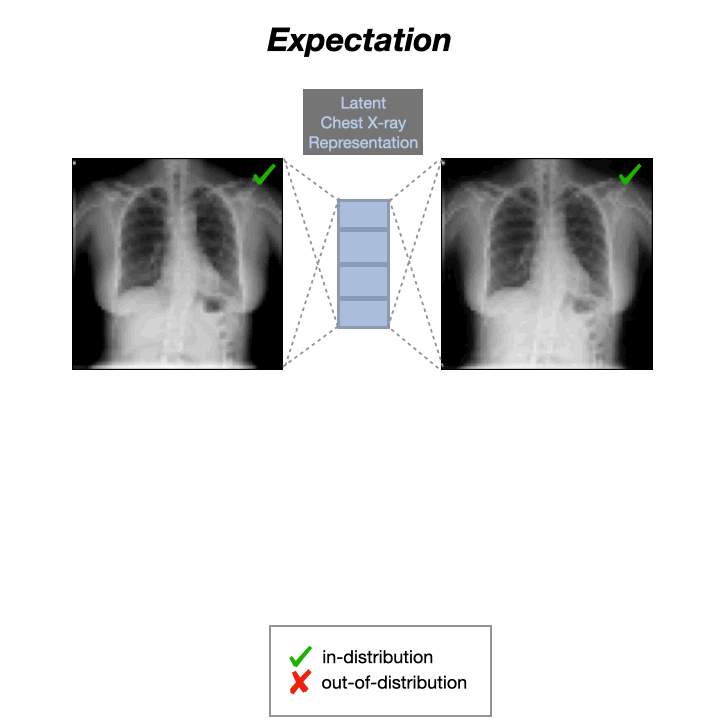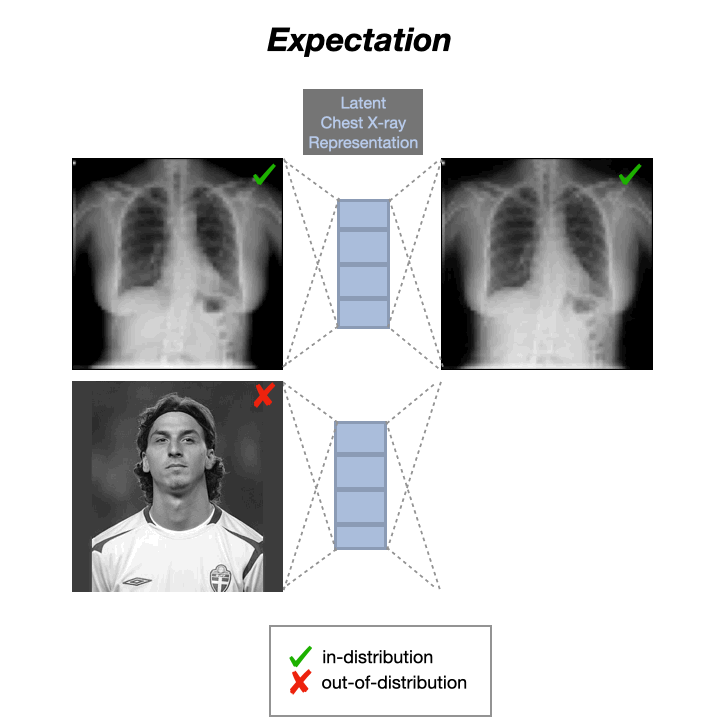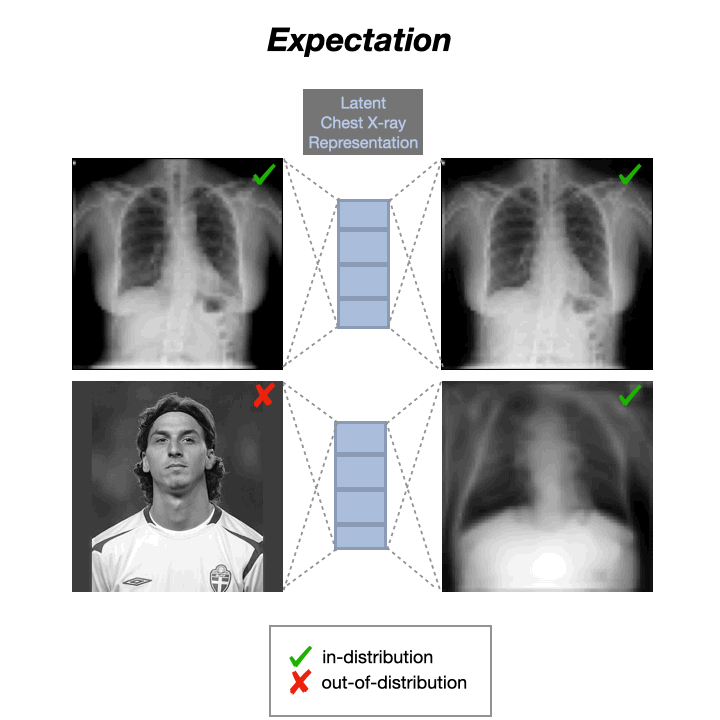
An anomaly score can thus be directly computed from the residual of the input $x$ and its reconstruction: $s(x) = |x-g(f(x))|$, with $f$ and $g$ being the encoding and decoding functions, usually implemented with neural networks.
Limitations:
However, recent literature has shown that AEs are able to reconstruct OoD samples sometimes even better than ID samples:

Main findings:
- OoD metrics might hide correlations such as, background intensties, or domain shifts.
- We propose two metrics based on the FID and classifier confidence to investigate the distribution learning capacity of state-of-the-art (SOTA) AEs
- SOTA AEs are either unable to constrain the latent manifold and allow reconstruction of abnormal patterns, or they are failing to accurately restore the inputs from their latent distribution, resulting in blurred or misaligned reconstructions.
- We propose novel deformable auto-encoders (MorphAEus) to learn perceptually aware global image priors and locally adapt their morphometry based on estimated dense deformation fields, drastically reducing the false positives.
MorphAEus: Deformable Auto-Encoders
To the best of our knowledge, this is the first work that uses dense deformations fields for unsupervised anomaly detection. We propose to learn minimal and sufficient representations for OoD by leveraging deep AEs to predict global image priors that match the training distribution and locally adapting their morphometry with estimated deep deformation fields to better match the input:
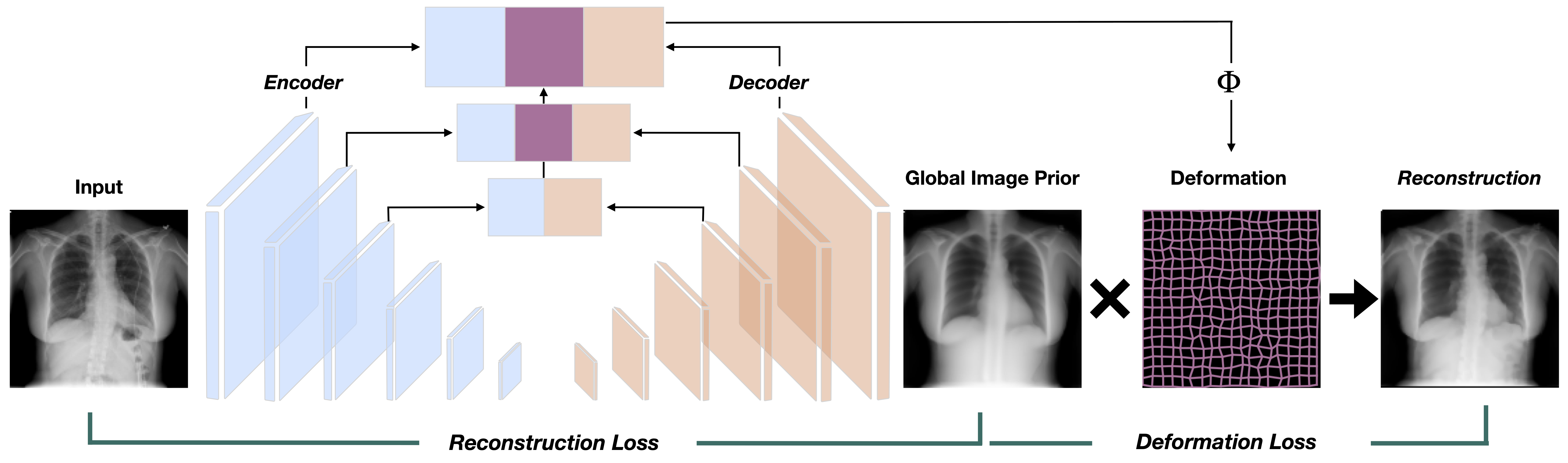
The intuition behind using deformation fields is to align healthy tissues to the input image, reducing the number of false positives, without altering the true positive pathology detection:
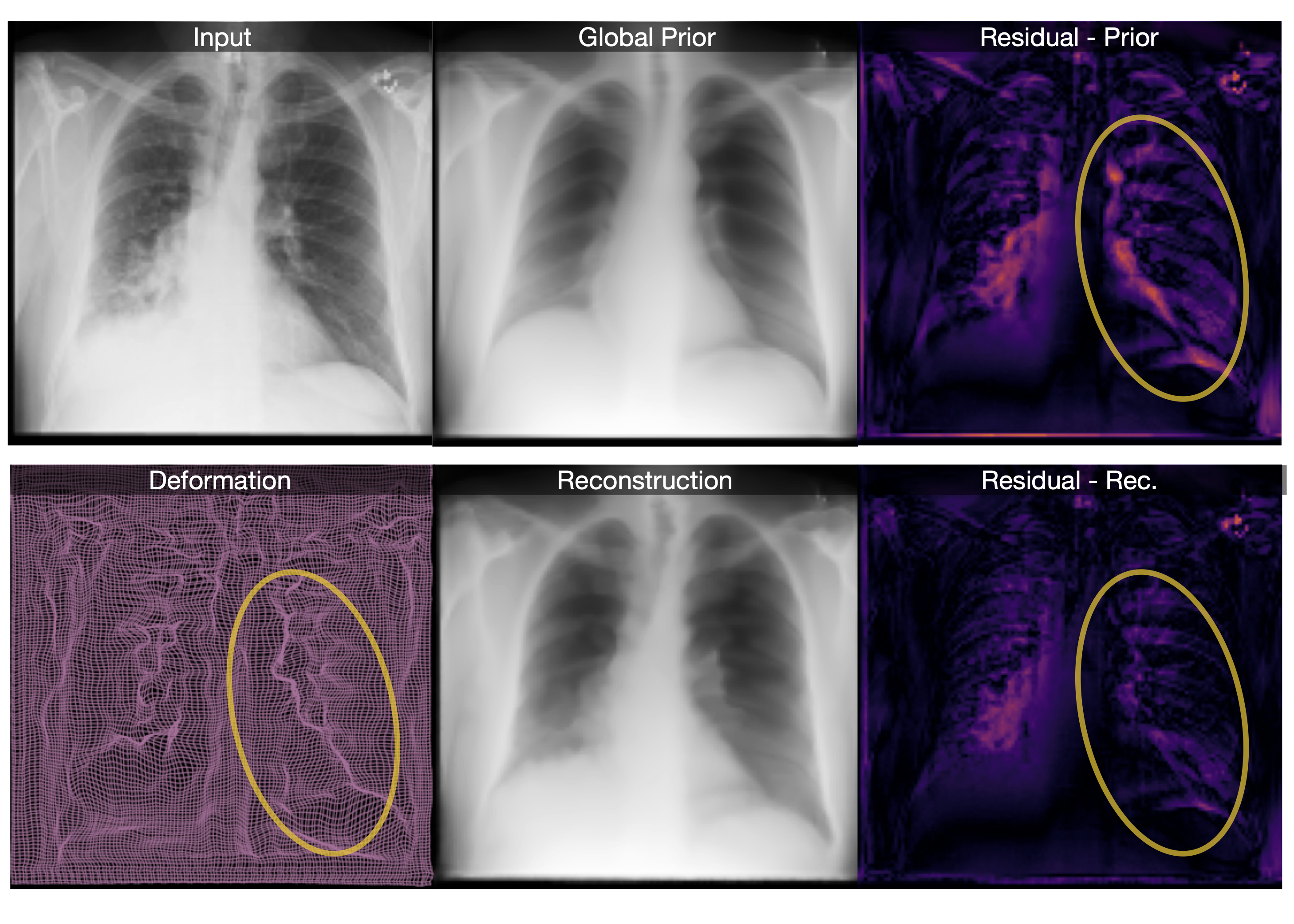
Our method reconstructs healthy synthesis of pathological inputs, enabling the localization of pathology:
Do we learn the manifold? On OoD detection
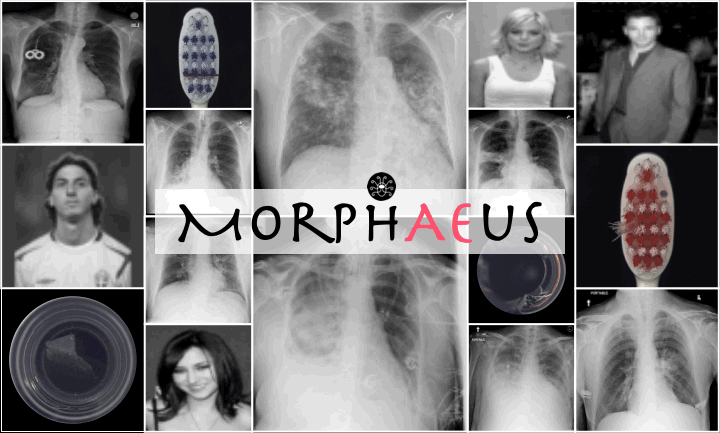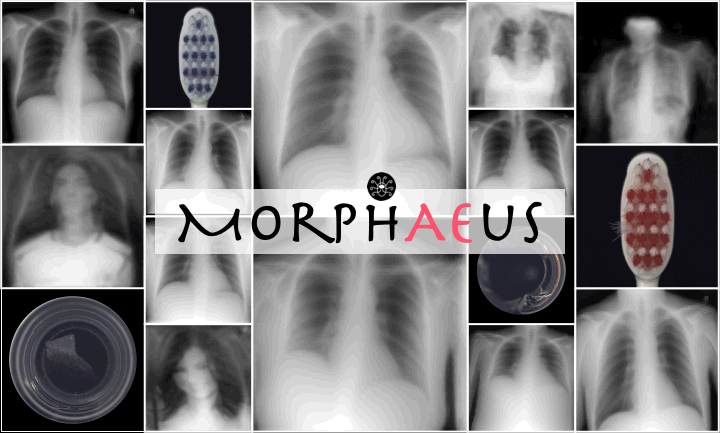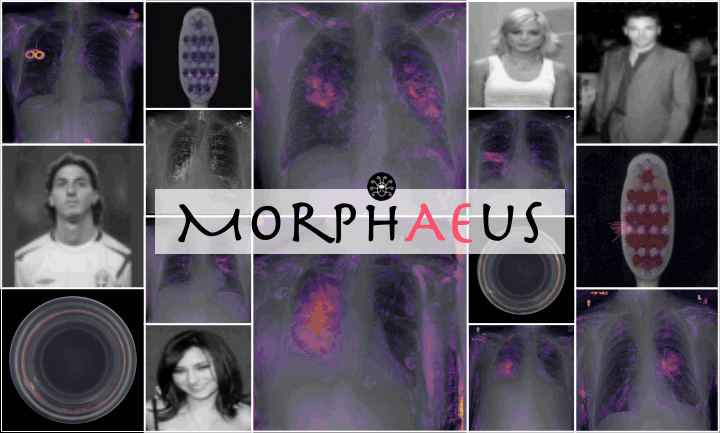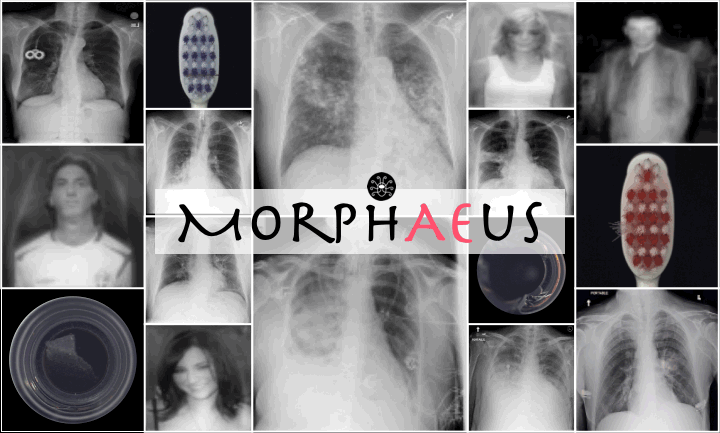
Reconstructions of ID and OoD samples by SOTA AEs:
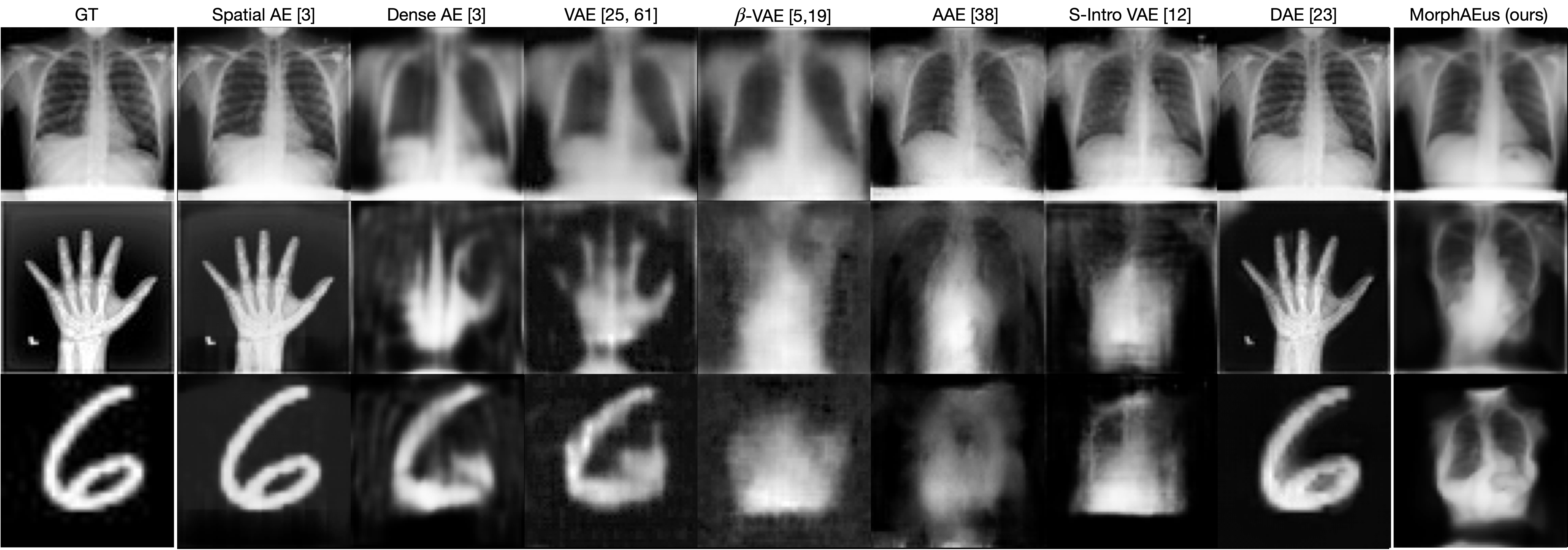
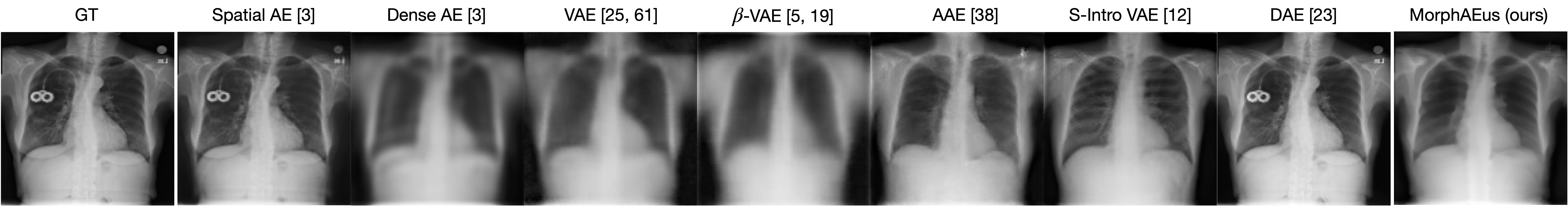
SOTA AEs either accurately reconstruct the input, but copy abnormalities (e.g., hand, and digits) or yield blury (Dense AE, VAE, $\beta$-VAE) or missaligned (adversarial AEs) reconstructions. Our proposed method learns the training distribution and produces detailed reconstructions of ID samples.
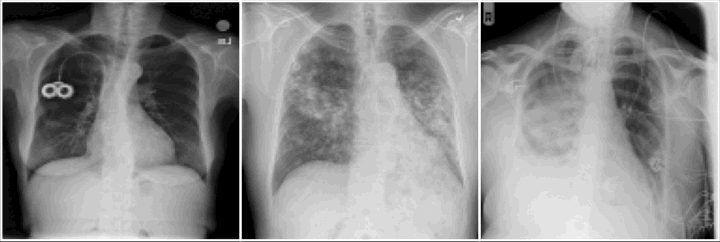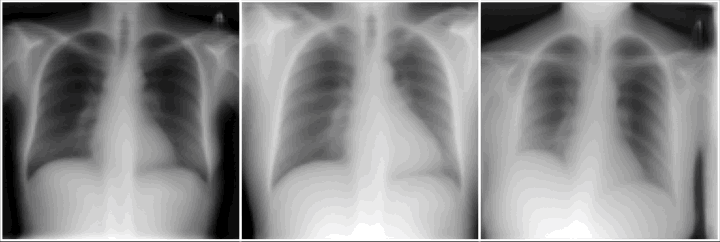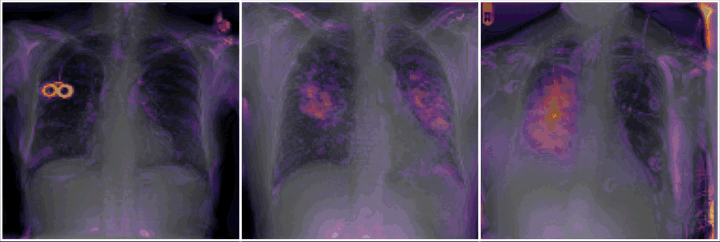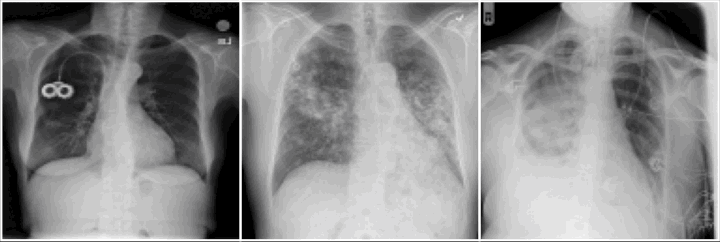
Do we learn the healthy manifold? On pathology detection
Healthy example:
Pathology examples:
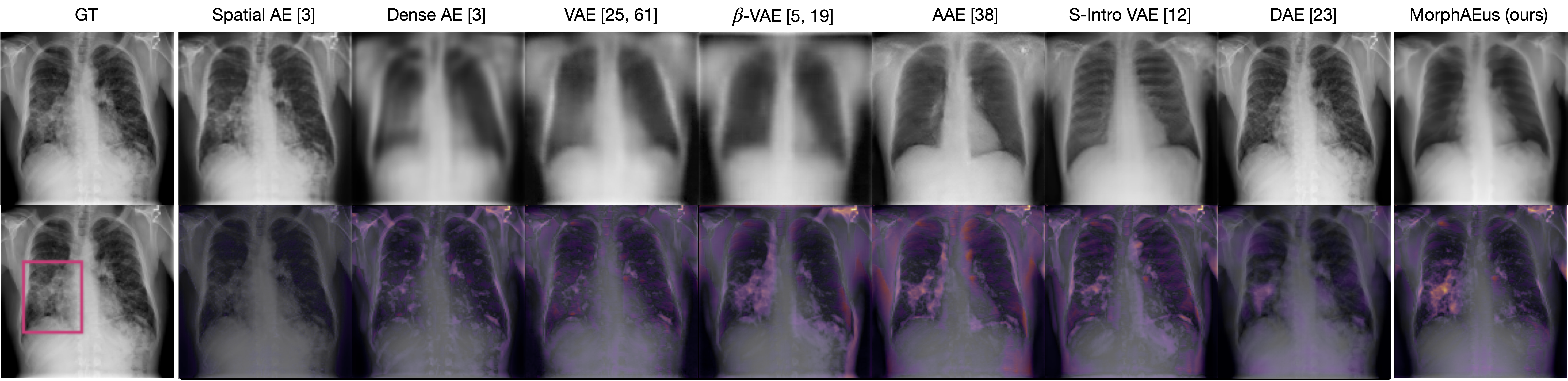
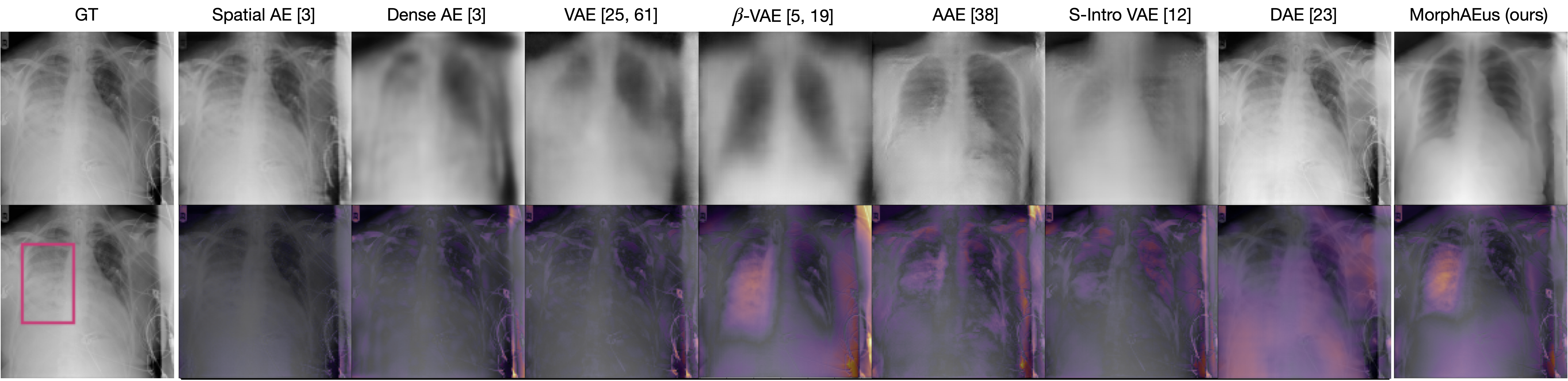
We synthesize detailed healthy reconstructions of pathological inputs, enabling unsupervised disease detection and localization.
Check our publication below for more information.